Users
Social media
- More details here...
- Address
Parc Científic de la Universitat de València C/
Catedrático Agustín Escardino, 9
46980 Paterna (Valencia) Spain - Email:
iu.i2sysbio@uv.es - Phone:
(+34) 963544810
- Address
Links
Deciphering ‘the tomato’s internal language’: how its genes communicate to resist drought and improve fruit quality

Investigation
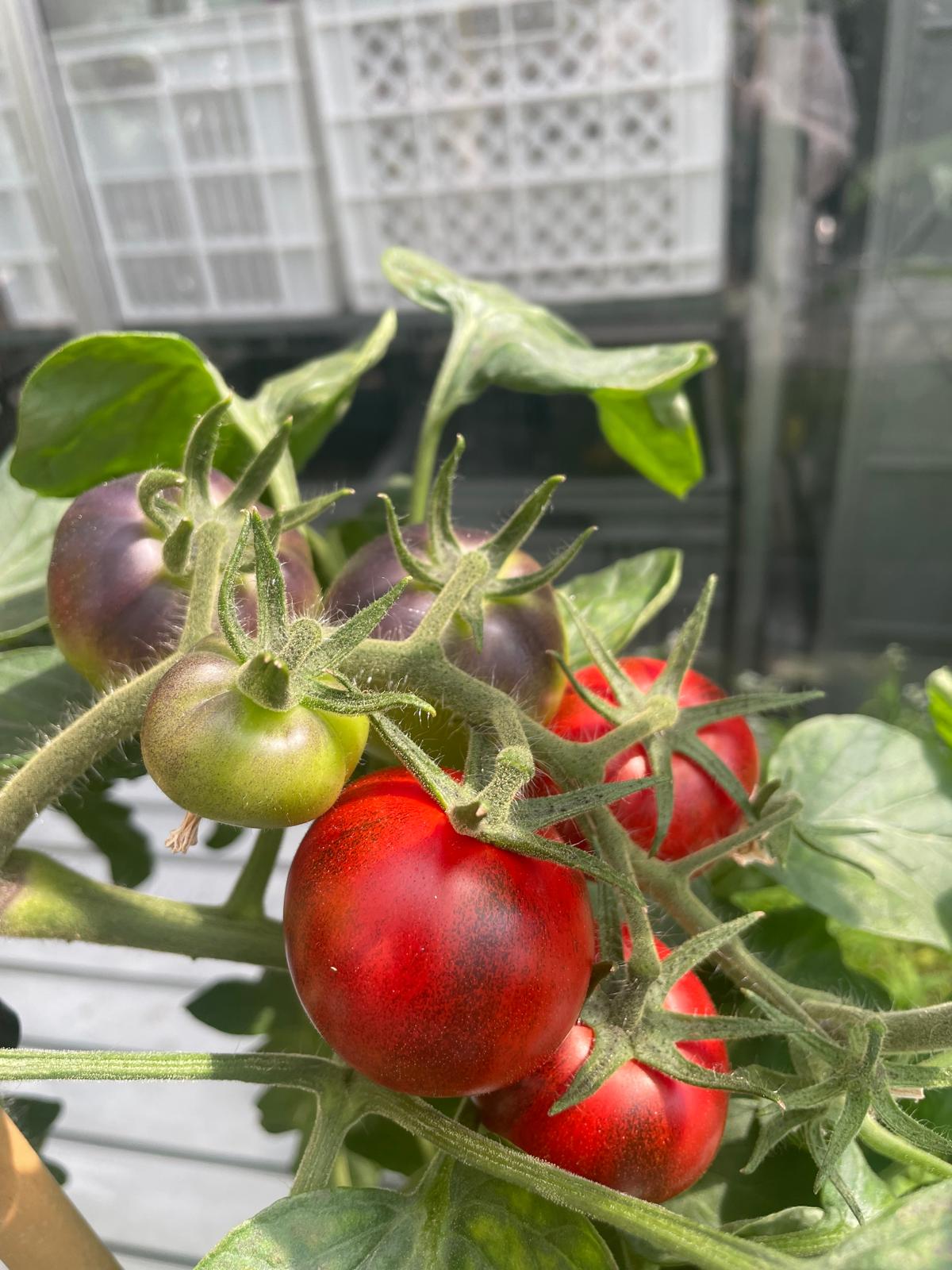
Deciphering ‘the tomato’s internal language’: how its genes communicate to resist drought and improve fruit quality
A study by the Institute of Integrative Systems Biology (I2SysBio, CSIC-UV) and the Phytolearning Millennium Nucleus (Chile) has deciphered how tomato (Solanum lycopersicum) genes communicate with each other to coordinate essential processes such as fruit ripening and drought response. This finding, published in the journal Plant Communications, opens new avenues for developing more resilient and sustainable crops in the context of climate change. The publication was also featured on the cover of the November 2025 issue.
The study, led by Dr Tomás Matus, researcher at I2SysBio, Dr Elena Vidal and Dr José Miguel Álvarez, both directors of Núcleo Milenio Phytolearning, reveals that the functioning of the tomato plant depends on complex interaction networks, where each organ – roots, leaves, flowers and fruits – organises its own regulatory strategy.
To achieve this, the team analysed more than 10,000 gene expression data sets from different organs and environmental conditions and reconstructed how genes communicate with each other. ‘What we ultimately achieved was to understand who gives the orders, who responds, and how that conversation changes between a root, a leaf or a fruit’, explains Vidal.
This work has also made it possible to generate a genuine ‘functional map’ of tomato metabolism, identifying the most influential nodes in the network: genes that act as coordinators of the response to water stress (drought) and fruit development. ‘With this information, we can design smarter genetic enhancement strategies based on complete networks rather than isolated hypotheses’, says Matus, co-author of the article and leader of the TomsBio Lab at I2SysBio.
A ‘network’ approach to climate change and drought
For decades, public discourse and research on how to improve crops has focused on finding the ‘miracle gene’. But this research marks a paradigm shift: modifying a single gene can have knock-on effects throughout the entire network, requiring strategies based on complete systems.
‘Adopting a “network” view allows us to understand that in plants there are no genes that act in isolation, but rather complex communication systems where each gene influences many others’, explains Matus. In contexts such as climate change and drought, this perspective is key because it helps us discover how plants reorganise their internal networks to adapt to stress: which genes take on leadership roles, how regulatory priorities change between roots, leaves or fruits, and which communication mechanisms are activated or deactivated.
‘Where crops face increasingly extreme conditions, understanding these networks can help us anticipate and select varieties with more efficient resilience strategies, rather than focusing on a single “miracle gene”. It is a more realistic and modern way of understanding plant biology in the face of climate change,’ says Matus.
TomViz: an open web platform
As part of the study, the team has created TomViz, an interactive platform that allows users to explore tomato gene regulatory networks in a simple and visual way. This tool, integrated into the PlantaeViz environment, offers the scientific community open access to data and functionalities to consult genes, identify their connections and generate customised sub-networks. It also includes options for performing enrichment analyses, visualising the position of genes in the genome and downloading results in different formats.
Thanks to TomViz, any researcher—in Chile, Spain, or anywhere in the world—can take advantage of this resource to propose new strategies that make crops more resistant to drought, productive, and sustainable, promoting global collaboration and innovation in genetic improvement.
Reference:
Fernández J.D., Navarro-Payá D., Santiago A., Cerda A., Canan J., Contreras-Riquelme S., Moyano T.C., Landaeta-Sepúlveda D., Melet L., Canales J., Johnson N.R., Álvarez J.M., Matus J.T., y Vidal E.A (2025). Organ-level gene-regulatory networks inferred from transcriptomic data reveal context-specific regulation and highlight novel regulators of ripening and ABA-mediated responses in tomato. Plant Communitacions, 2025 DOI: 10.1016/j.xplc.2025.101499